Pixel-Level Deep Segmentation: Artificial Intelligence Quantifies Muscle on Computed Tomography for Body Morphometric Analysis
https://link.springer.com/article/10.1007/s10278-017-9988-z/fulltext.html
Paper published titled, “Pixel-Level Deep Segmentation: Artificial Intelligence Quantifies Muscle on Computed Tomography for Body Morphometric Analysis”
Lee, H., Troschel, F.M., Tajmir, S. et al. J Digit Imaging (2017). doi:10.1007/s10278-017-9988-z.
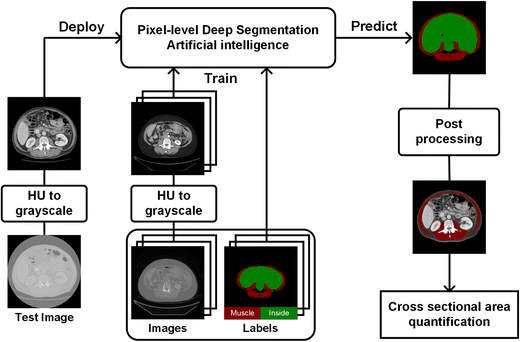
Pretreatment risk stratification is key for personalized medicine. While many physicians rely on an “eyeball test” to assess whether patients will tolerate major surgery or chemotherapy, “eyeballing” is inherently subjective and difficult to quantify. The concept of morphometric age derived from cross-sectional imaging has been found to correlate well with outcomes such as length of stay, morbidity, and mortality. However, the determination of the morphometric age is time intensive and requires highly trained experts. In this study, we propose a fully automated deep learning system for the segmentation of skeletal muscle cross-sectional area (CSA) on an axial computed tomography image taken at the third lumbar vertebra. We utilized a fully automated deep segmentation model derived from an extended implementation of a fully convolutional network with weight initialization of an ImageNet pre-trained model, followed by post processing to eliminate intramuscular fat for a more accurate analysis. This experiment was conducted by varying window level (WL), window width (WW), and bit resolutions in order to better understand the effects of the parameters on the model performance. Our best model, fine-tuned on 250 training images and ground truth labels, achieves 0.93 ± 0.02 Dice similarity coefficient (DSC) and 3.68 ± 2.29% difference between predicted and ground truth muscle CSA on 150 held-out test cases. Ultimately, the fully automated segmentation system can be embedded into the clinical environment to accelerate the quantification of muscle and expanded to volume analysis of 3D datasets.
